Entry
Reader's guide
Entries A-Z
Subject index
Metric Multidimensional Scaling
Metric multidimensional scaling (MDS) transforms a distance matrix into a set of coordinates such that the (Euclidean) distances derived from these coordinates approximate as well as possible the original distances. The basic idea of MDS is to transform the distance matrix into a cross-product matrix and then to find its eigendecomposition, which gives a principal component analysis (PCA). Like PCA, MDS can be used with supplementary or illustrative elements that are projected onto the dimensions after they have been computed.
An Example
The example is derived from O'Toole, Jiang, Abdi, and Haxby, who used a combination of principal component analysis and neural networks to analyze brain imaging data. In this study, 6 subjects were scanned using fMRI when they were watching pictures from 8 categories (faces, houses, cats, chairs, shoes, scissors, bottles, and scrambled images). The authors computed for each subject a distance matrix corresponding to how well they could predict the type of pictures that the subject was watching from his or her brain scans. The distance used was d′, which expresses the discriminability between categories.
O'Toole et al. give two distance matrices. The first one is the average distance matrix computed from the brain scans of all 6 subjects. The authors also give a distance matrix derived directly from the pictures watched by the subjects. The authors computed this distance matrix with the same algorithm that they used for the brain scans; they just substituted images for brain scans.
We will use these two matrices to review the basics of multidimensional scaling: namely, how to transform a distance matrix into a cross-product matrix and how to project a set of supplementary observations onto the space obtained by the original analysis.
Multidimensional Scaling: Eigenanalysis of a Distance Matrix
PCA is obtained by performing the eigendecomposition of a matrix. This matrix can be a correlation matrix (i.e., the variables to be analyzed are centered and normalized), a covariance matrix (i.e., the variables are centered but not normalized), or a cross-product matrix (i.e., the variables are neither centered nor normalized). A distance matrix cannot be analyzed directly using the eigendecomposition (because distance matrices are not positive semidefinite matrices), but it can be transformed into an equivalent cross-product matrix, which can then be analyzed.
Transforming a Distance Matrix into a Cross-Product Matrix
In order to transform a distance matrix into a crossproduct matrix, we start from the observation that the scalar product between two vectors can be transformed easily into a distance (the scalar product between vectors corresponds to a cross-product matrix). Let us start with some definitions. Suppose that a and b are two vectors with I elements. The Euclidean distance between these two vectors is computed as
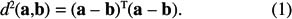
This distance can be rewritten to isolate the scalar product between vectors a and b:
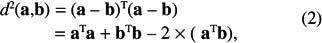
where aTb is the scalar product between a and b.
If the data are stored into an I × J data matrix denoted X (where I observations are described by J variables), the between-observations cross-product matrix is then obtained as

A distance matrix can be computed directly from the cross-product matrix
...
- Biographies
- Babbage, Charles
- Bernoulli, Jakob
- Bonferroni, Carlo Emilio
- Bruno, James Edward
- Comrey, Andrew L.
- Cronbach, Lee J.
- Darwin, Charles
- Deming, William Edwards
- Fisher, Ronald Aylmer
- Galton, Sir Francis
- Gauss, Carl Friedrich
- Gresham, Frank M.
- Jackson, Douglas N.
- Malthus, Thomas
- Markov, Andrei Andreevich
- Pascal, Blaise
- Pearson, Karl
- Poisson, Siméon Denis
- Reynolds, Cecil R.
- Torrance, E. Paul
- Wilcoxon, Frank
- Charts, Graphs, and Visual Displays
- Computer Topics and Tools
- Concepts and Issues in Measurement
- T Scores
- z Scores
- Ability Tests
- Achievement Tests
- Alternate Assessment
- Americans with Disabilities Act
- Anthropometry
- Aptitude Tests
- Artificial Neural Network
- Asymmetry of G
- Attitude Tests
- Basal Age
- Categorical Variable
- Classical Test Theory
- Coefficient Alpha
- Completion Items
- Computerized Adaptive Testing
- Construct Validity
- Content Validity
- Criterion Validity
- Criterion-Referenced Test
- Cronbach, Lee J.
- Curriculum-Based Measurement
- Diagnostic Validity
- Educational Testing Service
- Equivalence Testing
- Essay Items
- Ethical Issues in Testing
- Face Validity
- Gf-Gc Theory of Intelligence
- Guttman Scaling
- Health Insurance Portability and Accountability Act
- High-Stakes Tests
- Immediate and Delayed Memory Tasks
- Individuals with Disabilities Education Act
- Information Referenced Testing
- Informed Consent
- Intelligence Quotient
- Intelligence Tests
- Internal Review Board
- Interrater Reliability
- Interval Level of Measurement
- Ipsative Measure
- Item and Test Bias
- Item Response Theory
- KR-20 and KR-21
- Likert Scaling
- Measurement
- Measurement Error
- Metric Multidimensional Scaling
- Multiple-Choice Items
- Multitrait Multimethod Matrix and Construct Validity
- Nomothetic Versus Idiographic
- Ordinal Level of Measurement
- Parallel Forms Reliability
- Performance IQ
- Performance-Based Assessment
- Personality Tests
- Portfolio Assessment
- Predictive Validity
- Projective Testing
- Q Methodology
- Questionnaires
- Ratio Level of Measurement
- Reliability Theory
- Response to Intervention
- Reverse Scaling
- Scaling
- Section 504 of the Rehabilitation Act of 1973
- Self-Report
- Semantic Differential
- Semantic Differential Scale
- Six Sigma
- Spearman's Rho
- Split Half Reliability
- Standard Error of Measurement
- Standard Scores
- Standards for Educational and Psychological Testing
- Test-Retest Reliability
- Thurstone Scaling
- Torrance, E. Paul
- True/False Items
- Validity Coefficient
- Validity Theory
- Verbal IQ
- Concepts and Issues in Statistics
- Artificial Neural Network
- Attenuation, Correction for
- Autocorrelation
- Bayesian Statistics
- Bioinformatics
- Central Limit Theorem
- Decision Theory
- Diggle-Kenward Model for Dropout
- DISTATIS
- Exploratory Factor Analysis
- Factorial Design
- Fourier Transform
- Generalized Additive Model
- Generalized Method of Moments
- Generalized Procrustes Analysis
- Graphical Statistical Methods
- Hierarchical Linear Modeling
- Historiometrics
- Logistic Regression Analysis
- Loglinear Analysis
- Markov Chain Monte Carlo Methods
- Matrix Operations
- Mean
- Measurement Error
- Mixtures of Experts
- Nonparametric Statistics
- Propensity Scores
- Rasch Measurement Model
- Regression Analysis
- Sampling Distribution of a Statistic
- Signal Detection Theory
- Simpson's Paradox
- Spurious Correlation
- Standard Error of the Mean
- Standard Scores
- Support Vector Machines
- Survival Analysis
- Type I Error
- Type II Error
- Data and Data Reduction Techniques
- Descriptive Statistics
- Arithmetic Mean
- Attenuation, Correction for
- Autocorrelation
- Average
- Average Deviation
- Bayley Scales of Infant Development
- Biserial Correlation Coefficient
- Class Interval
- Coefficients of Correlation, Alienation, and Determination
- Cognitive Psychometric Assessment
- Cohen's Kappa
- Correlation Coefficient
- Cumulative Frequency Distribution
- Deviation Score
- Difference Score
- Estimates of the Population Median
- Fisher's Z Transformation
- Frequency Distribution
- Galton, Sir Francis
- Grand Mean
- Harmonic Mean
- Histogram
- Kendall Rank Correlation
- Mean
- Measures of Central Tendency
- Median
- Mode
- Moving Average
- Parameter
- Parameter Invariance
- Part and Partial Correlations
- Pearson Product-Moment Correlation Coefficient
- Pearson, Karl
- Percentile and Percentile Rank
- Scattergram
- Semi-Interquartile Range
- Spurious Correlation
- Standard Deviation
- Survey Weights
- Text Analysis
- Evaluation
- Experimental Methods
- Alternative Hypothesis
- American Statistical Association
- Americans with Disabilities Act
- Association for Psychological Science
- Basic Research
- Bioinformatics
- Complete Independence Hypothesis
- Continuous Variable
- Critical Value
- Data Collection
- Data Mining
- Delphi Technique
- Dependent Variable
- Descriptive Research
- Ethical Issues in Testing
- Ethical Principles in the Conduct of Research With Human Participants
- Fractional Randomized Block Design
- Hello-Goodbye Effect
- Hypothesis and Hypothesis Testing
- Independent Variable
- Informed Consent
- Instrumental Variables
- Internal Review Board
- Longitudinal/Repeated Measures Data
- Meta-Analysis
- Missing Data Method
- Mixed Models
- Mixture Models
- Moderator Variable
- Monte Carlo Methods
- Null Hypothesis Significance Testing
- Ockham's Razor
- Pairwise Comparisons
- Post Hoc Comparisons
- Projective Testing
- Quasi-Experimental Method
- Sample Size
- Section 504 of the Rehabilitation Act of 1973
- Significance Level
- Simple Main Effect
- Simulation Experiments
- Single-Subject Designs
- Standards for Educational and Psychological Testing
- Statistical Significance
- Suppressor Variable
- Variable
- Variable Deletion
- Variance
- Inferential Statistics
- Akaike Information Criterion
- Analysis of Covariance (ANCOVA)
- Analysis of Variance (ANOVA)
- Bayes Factors
- Bayesian Information Criterion
- Binomial Test
- Bonferroni, Carlo Emilio
- Complete Independence Hypothesis
- Data Analysis ToolPak
- Exploratory Factor Analysis
- Factorial Design
- Fisher, Ronald Aylmer
- Hierarchical Linear Modeling
- Hypothesis and Hypothesis Testing
- Inferential Statistics
- Logistic Regression Analysis
- Markov, Andrei Andreevich
- Null Hypothesis Significance Testing
- Pairwise Comparisons
- Part and Partial Correlations
- Repeated Measures Analysis of Variance
- Type I Error
- Type II Error
- Wilcoxon, Frank
- Organizations and Publications
- Abstracts
- American Doctoral Dissertations
- American Psychological Association
- American Statistical Association
- Association for Psychological Science
- Buros Institute of Mental Measurements
- Centers for Disease Control and Prevention
- Educational Testing Service
- Journal of Modern Applied Statistical Methods
- Journal of Statistics Education
- Journal of the American Statistical Association
- National Science Foundation
- Psychometrics
- PsycINFO
- Society for Research in Child Development
- Prediction and Estimation
- Attributable Risk
- Bernoulli, Jakob
- Chance
- Conditional Probability
- Confidence Intervals
- Continuous Variable
- Curse of Dimensionality
- Decision Boundary
- Decision Theory
- File Drawer Problem
- Gambler's Fallacy
- Generalized Estimating Equations
- Law of Large Numbers
- Maximum Likelihood Method
- Nonprobability Sampling
- Pascal, Blaise
- Probability Sampling
- Random Numbers
- Relative Risk
- Signal Detection Theory
- Significance Level
- Three-Card Method
- Probability
- Qualitative Methods
- Samples, Sampling, and Distributions
- Acceptance Sampling
- Adaptive Sampling Design
- Age Norms
- Attrition Bias
- Career Maturity Inventory
- Central Limit Theorem
- Class Interval
- Cluster Sampling
- Confidence Intervals
- Convenience Sampling
- Cumulative Frequency Distribution
- Data Collection
- Diggle-Kenward Model for Dropout
- Gauss, Carl Friedrich
- Heteroscedasticity and Homoscedasticity
- Homogeneity of Variance
- Hypergeometric Distribution
- Kurtosis
- Malthus, Thomas
- Multicollinearity
- Multivariate Normal Distribution
- Nonprobability Sampling
- Normal Curve
- Ogive
- Parameter
- Percentile and Percentile Rank
- Poisson Distribution
- Poisson, Siméon Denis
- Posterior Distribution
- Prior Distribution
- Probability Sampling
- Quota Sampling
- Random Sampling
- Sample
- Sample Size
- Semi-Interquartile Range
- Simpson's Rule
- Skewness
- Smoothing
- Stanine
- Stratified Random Sampling
- Unbiased Estimator
- Statistical Techniques
- k-Means Cluster Analysis
- t Test for Two Population Means
- Binomial Distribution/Binomial and Sign Tests
- Bivariate Distributions
- Bonferroni Test
- Bowker Procedure
- Causal Analysis
- Centroid
- Chance
- Chi-Square Test for Goodness of Fit
- Chi-Square Test for Independence
- Classification and Regression Tree
- Cochran Q Test
- Cohen's Kappa
- Delta Method
- Dimension Reduction
- Discriminant Analysis
- Dissimilarity Coefficient
- Dixon Test for Outliers
- Dunn's Multiple Comparison Test
- Eigendecomposition
- Eigenvalues
- EM Algorithm
- Exploratory Data Analysis
- Factor Analysis
- Factor Scores
- Fisher Exact Probability Test
- Fisher's LSD
- Friedman Test
- Goodness-of-Fit Tests
- Grounded Theory
- Kolmogorov-Smirnov Test for One Sample
- Kolmogorov-Smirnov Test for Two Samples
- Kruskal-Wallis One-Way Analysis of Variance
- Latent Class Analysis
- Likelihood Ratio Test
- Lilliefors Test for Normality
- Mann-Whitney U Test (Wilcoxon Rank-Sum Test)
- McNemar Test for Significance of Changes
- Median Test
- Meta-Analysis
- Multiple Comparisons
- Multiple Factor Analysis
- Multiple Imputation for Missing Data
- Multivariate Analysis of Variance (MANOVA)
- Newman-Keuls Test
- O'Brien Test for Homogeneity of Variance
- Observational Studies
- One-Way Analysis of Variance
- Page's L Test
- Paired Samples t Test (Dependent Samples t Test)
- Path Analysis
- Peritz Procedure
- Scan Statistic
- Shapiro-Wilk Test for Normality
- Structural Equation Modeling
- Tests of Mediating Effects
- Three-Card Method
- Tukey-Kramer Procedure
- Wilcoxon Signed Ranks Test
- Statistical Tests
- t Test for Two Population Means
- Analysis of Covariance (ANCOVA)
- Analysis of Variance (ANOVA)
- Behrens-Fisher Test
- Binomial Distribution/Binomial and Sign Tests
- Binomial Test
- Bonferroni Test
- Bowker Procedure
- Chi-Square Test for Goodness of Fit
- Chi-Square Test for Independence
- Classification and Regression Tree
- Cochran Q Test
- Dixon Test for Outliers
- Dunn's Multiple Comparison Test
- Excel Spreadsheet Functions
- Fisher Exact Probability Test
- Fisher's LSD
- Friedman Test
- Goodness-of-Fit Tests
- Kolmogorov-Smirnov Test for One Sample
- Kolmogorov-Smirnov Test for Two Samples
- Kruskal-Wallis One-Way Analysis of Variance
- Latent Class Analysis
- Likelihood Ratio Test
- Lilliefors Test for Normality
- Mann-Whitney U Test (Wilcoxon Rank-Sum Test)
- McNemar Test for Significance of Changes
- Median Test
- Multiple Comparisons
- Multivariate Analysis of Variance (MANOVA)
- Newman-Keuls Test
- O'Brien Test for Homogeneity of Variance
- One- and Two-Tailed Tests
- One-Way Analysis of Variance
- Page's L Test
- Paired Samples t Test (Dependent Samples t Test)
- Peritz Procedure
- Repeated Measures Analysis of Variance
- Shapiro-Wilk Test for Normality
- Tests of Mediating Effects
- Tukey-Kramer Procedure
- Wilcoxon Signed Ranks Test
- Tests by Name
- Adjective Checklist
- Alcohol Use Inventory
- Armed Forces Qualification Test
- Armed Services Vocational Aptitude Battery
- Basic Personality Inventory
- Bayley Scales of Infant Development
- Beck Depression Inventory
- Behavior Assessment System for Children
- Bender Visual Motor Gestalt Test
- Bracken Basic Concept Scale–Revised
- California Psychological Inventory
- Career Assessment Inventory
- Career Development Inventory
- Career Maturity Inventory
- Carroll Depression Scale
- Children's Academic Intrinsic Motivation Inventory
- Clinical Assessment of Attention Deficit
- Clinical Assessment of Behavior
- Clinical Assessment of Depression
- Cognitive Abilities Test
- Cognitive Psychometric Assessment
- Comrey Personality Scales
- Coping Resources Inventory for Stress
- Culture Fair Intelligence Test
- Differential Aptitude Test
- Ecological Momentary Assessment
- Edwards Personal Preference Schedule
- Embedded Figures Test
- Fagan Test of Infant Intelligence
- Family Environment Scale
- Gerontological Apperception Test
- Goodenough Harris Drawing Test
- Graduate Record Examinations
- Holden Psychological Screening Inventory
- Illinois Test of Psycholinguistic Abilities
- Information Systems Interaction Readiness
- Internal External Locus of Control Scale
- International Assessment of Educational Progress
- Iowa Tests of Basic Skills
- Iowa Tests of Educational Development
- Jackson Personality Inventory–Revised
- Jackson Vocational Interest Survey
- Kaufman Assessment Battery for Children
- Kinetic Family Drawing Test
- Kingston Standardized Cognitive Assessment
- Kuder Occupational Interest Survey
- Laboratory Behavioral Measures of Impulsivity
- Law School Admissions Test
- Life Values Inventory
- Luria Nebraska Neuropsychological Battery
- Male Role Norms Inventory
- Matrix Analogies Test
- Millon Behavioral Medicine Diagnostic
- Millon Clinical Multiaxial Inventory-III
- Minnesota Clerical Test
- Minnesota Multiphasic Personality Inventory
- Multidimensional Aptitude Battery
- Multiple Affect Adjective Checklist–Revised
- Myers-Briggs Type Indicator
- NEO Personality Inventory
- Neonatal Behavioral Assessment Scale
- Peabody Picture Vocabulary Test
- Personal Projects Analysis
- Personality Assessment Inventory
- Personality Research Form
- Piers-Harris Children's Self-Concept Scale
- Preschool Language Assessment Instrument
- Profile Analysis
- Projective Hand Test
- Quality of Well-Being Scale
- Raven's Progressive Matrices
- Roberts Apperception Test for Children
- Rorschach Inkblot Test
- Sixteen Personality Factor Questionnaire
- Social Climate Scales
- Social Skills Rating System
- Spatial Learning Ability Test
- Stanford Achievement Test
- Stanford-Binet Intelligence Scales
- Strong Interest Inventory
- Stroop Color and Word Test
- Structured Clinical Interview for DSM-IV
- System of Multicultural Pluralistic Assessment
- Thematic Apperception Test
- Torrance Tests of Creative Thinking
- Torrance Thinking Creatively in Action and Movement
- Universal Nonverbal Intelligence Test
- Vineland Adaptive Behavior Scales
- Vineland Social Maturity Scale
- Wechsler Adult Intelligence Scale
- Wechsler Individual Achievement Test
- Wechsler Preschool and Primary Scale of Intelligence
- West Haven-Yale Multidimensional Pain Inventory
- Woodcock Johnson Psychoeducational Battery
- Woodcock Reading Mastery Tests Revised
- Loading...
Get a 30 day FREE TRIAL
-
Watch videos from a variety of sources bringing classroom topics to life
-
Read modern, diverse business cases
-
Explore hundreds of books and reference titles

Sage Recommends
We found other relevant content for you on other Sage platforms.

Have you created a personal profile? Login or create a profile so that you can save clips, playlists and searches