Entry
Reader's guide
Entries A-Z
Subject index
Intraclass Correlation
An intraclass correlation coefficient is a measure of association based on a variance ratio. Formally, it is the ratio of the variance for an object of measurement (σ2Object) to the object variance plus error variance:
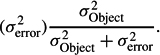
It differs from an analog measure of association known as OMEGA SQUARED (ω2) only in the statistical model used to develop the sample estimate of σ2Object. Intraclass correlation estimates are made using the random effects model (i.e., the objects in any one sample are randomly sampled from a population of potential objects) and ω2 estimates are made using the FIXED EFFECTS MODEL (i.e., the objects in the sample are the only ones of interest and could not be replaced without altering the question being addressed in the research).
Intraclass correlation is best introduced using one of the problems it was developed to solve—what the correlation of siblings is on a biometric trait. If the sample used to obtain the correlation estimate consisted just of sibling pairs (in which sibships are the object of measurement), a Pearson product-moment, interclass correlation could be used, but this procedure would not lead to a unique estimate. The reason is seen with an example. For three sibling pairs {1, 2}, {3, 4}, and {5, 6}, the Pearson r for pairs when listed in this original order {1, 2}{3, 4}{5, 6} is different from the Pearson r for the same pairs if listed as {2, 1}{3, 4}{6, 5}, even though the sibling data are exactly the same. The Pearson procedure requires the totally arbitrary assignment of pair members to a set X and a set Y when, in fact, the only assignment is to sibship. How, then, is one to obtain a measure of correlation within sibship classes without affecting the estimate through arbitrary assignments? An early answer was to double-list each pair and then compute a Pearson r. For the example above, this meant computing the Pearson r for the twice-listed pairs {1, 2}{2, 1}{3, 4}{4, 3}{5, 6}{6, 5}.
Double-listing the data for each sibship leads to a convenient solution in the case of sibling pairs, but the method becomes increasingly laborious as sibships are expanded to include 3, 4, or k siblings. A computational advance was introduced by J. A. Harris (1913), and then R. A. Fisher (1925) applied his analysis of variance to the problem. This latter approach revealed that a sibling correlation is an application of the one-way random effects variance model. Inthismodel, each sibling value is conceptualized as the sum of three independent components: a population mean (μ) for the biometric trait in question; an effect for the sibship (α), which is a deviation of the sibship mean from the population mean; and a deviation of the individual sibling from the sibship mean (ε). In its conventional form, the model is written Yij = μ + αi + εij. The sibling correlation is then the variance of the alpha effects over the variance of the alpha effects plus epsilon (error) effects:
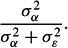
In other words, it is the proportion of total variance in the trait that is due to differences between sibships. For traits that run in families, the correlation would be high; for those that don't, the correlation would be low.
...
- Analysis of Variance
- Association and Correlation
- Association
- Association Model
- Asymmetric Measures
- Biserial Correlation
- Canonical Correlation Analysis
- Correlation
- Correspondence Analysis
- Intraclass Correlation
- Multiple Correlation
- Part Correlation
- Partial Correlation
- Pearson's Correlation Coefficient
- Semipartial Correlation
- Simple Correlation (Regression)
- Spearman Correlation Coefficient
- Strength of Association
- Symmetric Measures
- Basic Qualitative Research
- Basic Statistics
- F Ratio
- N(n)
- t-Test
- X¯
- Y Variable
- z-Test
- Alternative Hypothesis
- Average
- Bar Graph
- Bell-Shaped Curve
- Bimodal
- Case
- Causal Modeling
- Cell
- Covariance
- Cumulative Frequency Polygon
- Data
- Dependent Variable
- Dispersion
- Exploratory Data Analysis
- Frequency Distribution
- Histogram
- Hypothesis
- Independent Variable
- Measures of Central Tendency
- Median
- Null Hypothesis
- Pie Chart
- Regression
- Standard Deviation
- Statistic
- Causal Modeling
- Discourse/Conversation Analysis
- Econometrics
- Epistemology
- Ethnography
- Evaluation
- Event History Analysis
- Experimental Design
- Factor Analysis and Related Techniques
- Feminist Methodology
- Generalized Linear Models
- Historical/Comparative
- Interviewing in Qualitative Research
- Latent Variable Model
- Life History/Biography
- Log-Linear Models (Categorical Dependent Variables)
- Longitudinal Analysis
- Mathematics and Formal Models
- Measurement Level
- Measurement Testing and Classification
- Multilevel Analysis
- Multiple Regression
- Qualitative Data Analysis
- Sampling in Qualitative Research
- Sampling in Surveys
- Scaling
- Significance Testing
- Simple Regression
- Survey Design
- Time Series
- ARIMA
- Box-Jenkins Modeling
- Cointegration
- Detrending
- Durbin-Watson Statistic
- Error Correction Models
- Forecasting
- Granger Causality
- Interrupted Time-Series Design
- Intervention Analysis
- Lag Structure
- Moving Average
- Periodicity
- Serial Correlation
- Spectral Analysis
- Time-Series Cross-Section (TSCS) Models
- Time-Series Data (Analysis/Design)
- Trend Analysis
- Loading...
Get a 30 day FREE TRIAL
-
Watch videos from a variety of sources bringing classroom topics to life
-
Read modern, diverse business cases
-
Explore hundreds of books and reference titles

Sage Recommends
We found other relevant content for you on other Sage platforms.

Have you created a personal profile? Login or create a profile so that you can save clips, playlists and searches