Entry
Reader's guide
Entries A-Z
Subject index
SNP Technologies
A single nucleotide polymorphism (SNP or “snip”) is a single base pair variation at a particular location in the deoxyribonucleic acid (DNA) sequence that occurs in more than 1 percent of the population. SNPs are used as markers to identify regions in the genome associated with disease states such as obesity, diabetes, heart disease, and Alzheimer's. The discovery of such markers requires that a high number of SNPs be analyzed within large study populations. Many different technologies have been developed for SNP analysis that are rapid, accurate, and cost effective to increase the number of SNP markers suitable for use in determining effective medical care.
The human genome is composed of 3 billion base pairs of DNA, approximately 99.9 percent of which is identical between individuals. The 0.1 percent of genetic variation that occurs between us is responsible for differences in our appearance, personalities, and physiologies. These variations in DNA sequence, or “polymorphisms,” occur as deletions, insertions, sequence repeats, or SNPs. Mutations differ from polymorphisms in that they occur in less than 1 percent of the population.
SNPs account for approximately 90 percent of polymorphisms in the human genome, and over 9 million have been identified. Some SNPs are directly responsible for a disease process or for how an individual responds to a drug. When found within the coding region of a gene, a SNP may cause changes in the structure and possibly the function of the protein encoded by that gene. This could result from either a direct amino acid substitution or by affecting the splicing of the messenger ribonucleic acid (mRNA). If a SNP occurs in the regulatory region of a gene, the expression pattern of the gene could be significantly altered, with possible serious consequences to the cell or the organism.
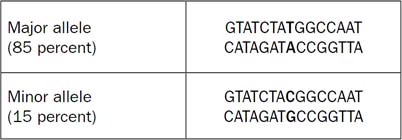
Figure 1. Example of Two Alleles Represented by an SNP
Most SNPs, however, are found in the 98 percent of the human genome that does not encode protein. Even when a SNP has no obvious affect on the expression of a gene, it may still be useful as a genetic or physical marker due to its proximity to a specific gene of interest. This is the basis for using SNPs to identify genes that either cause or influence various medical conditions, or to determine if an individual has a particular gene that predisposes him or her to a disease.
To understand how this works, it is first necessary to define a few terms. An allele is an alternative DNA sequence that occurs at a specific genetic locus. Figure 1 gives an example of two alleles represented by an SNP. At this location within the genome, a T-A base pair occurs 85 percent of the time, and a C-G base pair 15 percent in a given population. Most SNPs are biallelic, meaning only two sequence variations are found at that location. A few are known to be tri- or tetraallelic.
A particular combination of alleles on a given region of a chromosome is called a haplotype, whereas a collection of alleles within the entire genome is called a genotype.Figure 2 illustrates two different haplotypes, in which two SNPs are located near one another on a given chromosome.
...
- Biological or Genetic Contributors to Obesity
- Adipocytes
- Adiponectin
- Adrenergic Receptors
- Agouti and Agouti Related Protein
- Animal Models of Obesity
- Animal QTLs (Quantitative Trait Locus)
- Bardet-Biedl Syndromes
- Cannabinoid Receptor
- CD36 and FAT (Fatty Acid Transporters)
- Cholecystokinin (CCK)
- Cortisol
- Cushing Syndrome
- Cytokines
- Db/Db Mouse
- Dopamine Receptor
- Down's Syndrome
- Epistatic Effects of Genes on Obesity
- Estrogen-Related Receptor
- Familial Lipodystrophies
- Fatty Acid Transport Proteins
- G-Protein Coupled Receptors
- Genetic Taste Factors
- Ghrelin
- Glucagon Receptor
- Glucocorticoids
- Glucokinase
- Growth Hormone
- HDL Receptors
- Histamine Receptor
- Hormone Sensitive Lipase
- Human QTLs
- Hypothyroidism
- Insulin and Insulin Resistance
- Insulin-Like Growth Factors
- Interleukins
- Intrauterine Growth Restriction
- LDL Receptors
- Leptin
- Leptin Receptors
- Lipoprotein Lipase
- Low Birth Weight
- Melanocortins
- Mendelian Disorders Related to Obesity
- Metabolic Rate
- Monogenic Effects that Result in Obesity
- Neuropeptides
- NPY (Neuropeptide Y)
- Ob/Ob Mouse
- Obesity and the Immune System
- Obesity Gene Map
- Opioid Receptor
- Perilipins
- POMC (Proopiomelanocortin)
- PPAR (Peroxisome Proliferator-Activated Receptors)
- Prader-Willi Syndrome
- Protein Kinase
- Set or Settling Point
- Steroids
- Thrifty Gene Hypothesis
- Thrifty Gene Hypothesis and Obesity
- Thyroid Hormone
- TNF (Tumor Nucrosis Factors)
- Transgenics and Knockouts for Obesity-Related Genes
- Tubby Candidate Gene
- Twin Studies and Genetics of Obesity
- Uncoupling Proteins
- Viral Causes of Obesity
- Children and Obesity
- Advertising
- Atherosclerosis in Children
- Bariatric Surgery in Children
- Behavioral Treatment of Child Obesity
- Beverage Choices in Children and Obesity
- Breastfeeding
- Changing Children's Food Habits
- Childhood Obesity as a Risk Factor for Adult Overweight
- Childhood Obesity Treatment Centers
- Children and Diets
- Ethnic Disparities in the Prevalence of Childhood Obesity
- Family Behavioral Interventions
- Family Therapy in the Treatment of Overweight Children
- Flavor Programming and Childhood Food Preferences
- Food Intake Assessments in Children
- Formation and Development of Food Preferences
- Genetic Taste Factors
- Hypertension in Children
- Implications of Restriction of Foods on Child Feeding Habits
- Medical Interventions for Children
- Metabolic Disorders and Childhood Obesity
- Morbid Obesity in Children
- National Weight Loss Efforts for Children
- Overweight Children and School Performance
- Overweight Children and the Media
- Peer Influences on Obesity in Children
- Pharmacological Treatment of Childhood Obesity
- Physical Activity in Children
- Prevalence of Childhood Obesity in Developing Countries
- Prevalence of Childhood Obesity in the United States
- Prevalence of Childhood Obesity Worldwide
- Prevention
- School-Based Interventions to Prevent Obesity
- Self-Esteem and Children's Weight
- Stigmas against Overweight Children
- Type 2 Diabetes
- Dietary Interventions to Treat Obesity
- Atkins Diet
- Calcium and Dairy Products
- Caloric Restriction
- Carbohydrate “Addictions”
- Chromium Picolinate
- Diet Myths
- Dietary Restraint
- Exercise
- Fast Foods
- Fiber and Obesity
- Fruits and Vegetables
- High-Carbohydrate Diets
- High-Protein Diets
- Jenny Craig
- L.A. Weight Loss
- Liquid Diets
- Low-Calorie Diets
- Low-Fat Diets
- Macrodiets
- Medifast
- Non-Diet Approaches
- Nutrisystem
- Nutrition Fads
- Optifast
- Physical Activity and Obesity
- Portion Control
- Slim-Fast
- South Beach Diet
- Supplements and Obesity
- Vegetarianism
- Very Low-Calorie Diets
- Volumetrics
- Water and Obesity
- Weight Watchers
- Zone, The
- Disordered Eating and Obesity
- Anorexia Nervosa
- Antidepressants
- Appetite Signals
- Binge Eating
- Body Dysmorphic Disorder
- Body Image
- Bulimia Nervosa
- Childhood Onset Eating Disorders
- Cognitive-Behavioral Therapy
- Depression
- Dieting: Good or Bad?
- Disinhibited Eating
- DSM-IV
- Eating Disorders and Athletes
- Eating Disorders and Gender
- Eating Disorders and Obesity
- Eating Disorders in School Children
- EDNOS
- Families of Eating Disorder Patients
- Feminist Perspective and Body Image Disorders
- Genetic Influences on Eating Disorders
- Hunger
- Neurotransmitters
- Night Eating Syndrome
- Physiological Aspects of Anorexia
- Physiological Aspects of Bulimia
- Prevalence of Disordered Eating
- Sexual Abuse and Eating Disorders
- Treatment Centers for Eating Disorders
- Weight Cycling and Yo-Yo Dieting
- Environmental Contributors to Obesity
- Accessibility of Foods
- Advertising of Foods to Children
- Children's Television Programming
- Economics of Food
- Energy Density
- Fast Foods
- Food Advertising
- Food Labeling
- Governmental Subsidizing of Energy Dense Foods
- Inaccessibility of Exercise
- Increased Reliance on Automobiles
- Increasing Portion Sizes
- Palatability
- Parental and Home Environments
- Safe Play Opportunities for Children
- School Lunch Programs
- Schools and Obesity
- Sodas and Soft Drinks
- Sugar and Fat Substitutes
- Supersizing
- Television
- Toxic Environment
- Health Implications of Obesity
- Appetite Control
- Asthma
- Atherosclerosis
- Back Pain
- Blood Lipids
- Body Image
- Breast Cancer
- Colon Cancer
- Congestive Heart Failure
- Depression
- Elevated Cholesterol
- Fatty Liver
- Fertility
- Fitness
- Gallbladder Disease
- Gastroesophageal Reflux (GERD)
- Gastrointestinal Disorders
- Gestational Diabetes
- Gout
- High-Density Lipoproteins
- Hormones
- Hypertension
- Impotence
- Kidney Failure
- Kidney Stones
- Low-Density Lipoproteins
- Menstrual Problems
- Mortality and Obesity
- Osteoarthritis
- Osteoporosis
- Ovarian Cancer
- Ovarian Cysts
- Overall Diet Quality
- Polycystic Ovary Disease
- Respiratory Problems
- Sexual Health
- Sleep Apnea
- Stroke
- Type 2 Diabetes
- Urinary Incontinence in Severe Obesity in Women
- Uterine Cancers
- Medical Treatments for Obesity
- American Medical Association
- American Obesity Association
- Amphetamines
- Caffeine
- Cost of Medical Obesity Treatments
- Dexatrim
- Dieting: Good or Bad?
- Ephedra
- Fenfluramine
- Future of Medical Treatments for Obesity
- Gastric Bypass
- Gastroplasty
- Health Coverage of Gastric Surgeries
- International Obesity Task Force
- Laparoscopy
- Liquid Diets
- Low-Calorie Diets
- Medical Interventions for Children
- Medications that Affect Nutrient Partitioning
- Multidisciplinary Bariatric Programs
- Noradrenergic Drugs
- North American Association for the Study of Obesity
- Orlistat (Xenical)
- Physician-Assisted Weight Loss
- Qualifications for Gastric Surgery
- Roux-en-y Gastric Bypass
- Serotonergic Medications
- Sibutramine (Meridia)
- Thyroid Medications
- Vertical Banded Gastroplasty
- Very Low-Calorie Diets
- New Research Frontiers on Obesity
- Acomplia
- Bioelectrical Impedance Analysis
- Bod Pod and Pea Pod
- CART Peptides
- Combined Approaches to Treatment
- Computerized Tomography
- DEXA (Dual Energy X-ray Absorptiometry)
- Dilution Techniques
- Doubly Labeled Water
- Drug Targets that Decrease Food Intake/Appetite
- Drugs that Block Fat Cell Formation
- Energy Expenditure Technologies
- Food Technology
- Frontiers in Maintenance and Prevention
- Functional Foods
- Functional Magnetic Resonance Imaging
- Genetic Mapping of Obesity-Related Genes
- Genomics
- Histamines
- Hormone Disorders
- Hydrodensitrometry
- Indirect Calorimetry
- Intestinal Microflora Concentrations
- Leptin Supplements
- Magnetic Resonance Imaging Scans for Viewing Body Composition
- Metformin
- Microarray Analysis
- New Candidate Obesity Genes
- New Drug Targets that Prevent Fat Absorption
- New Drug Targets to Improve Insulin Sensitivity
- New Drug Targets to Increase Metabolic Rate
- Non-Diet Approaches
- Obesity and Viruses
- Quantitative Trait Locus Mapping
- Rimonabant
- SNP Technologies
- Three-D Image Reconstruction
- Translational Research
- Whole-Body Potassium Counting
- Obesity and Ethnicity/Race
- African Americans
- Asian Americans
- Body Fat Distribution in African Americans
- Body Fat Distribution in Asian Americans
- Body Fat Distribution in Hispanic Americans
- Cardiovascular Disease in African Americans
- Cardiovascular Disease in Asian Americans
- Cardiovascular Disease in Hispanic Americans
- Caucasians
- Dominican Americans
- Ethnic Variations in Body Fat Storage
- Ethnic Variations in Obesity-Related Health Risks
- Genetics
- Health Disparities—NIH Strategic Plan
- Hispanic Americans
- Hypertension in African Americans
- Hypertension in Asian Americans
- Hypertension in Hispanic Americans
- Mexican Americans
- Native Americans
- Obesity and Socioeconomic Status
- Pima Indians
- Puerto Rican Americans
- Sisters Together
- Thrifty Gene Hypothesis
- U.S. Office of Minority Health
- Western Diets
- Obesity and the Brain or Obesity and Behavior
- Antidepressants
- Appetite Control
- Autonomic Nervous System
- Bombesin
- Cannabinoid System
- Central Nervous System
- Cholecystokinin
- Conditioned Food Preferences
- Corticotropin-Releasing Hormone
- Dopamine
- Drugs and Food
- Fat Taste
- Flavor: Taste and Smell
- Folic Acid and Neural Tube Defects
- Food “Addictions”
- Food Reward
- Gustatory System
- Habituation
- Hypothalamus
- Inherited Taste Preferences
- Insulin
- Liking vs. Wanting
- Medications that Increase Body Weight
- Mood and Food
- Neuropeptide-Y
- Neurotransmitters
- Norepinephrine
- Nutrient Reward
- Olfactory System
- Opioids
- Oxytocin and Food Intake
- Peripheral Nervous Sytem
- Pituitary Gland
- Satietin
- Sensory-Specific Satiety
- Sweet Taste
- Sympathetic Nervous System
- Taste Aversion Learning
- Taste Reactivity
- Thyroid Gland
- Tryptophan
- Obesity as a Public Health Crisis
- Access to Nutritious Foods
- American Academy of Pediatrics
- American College of Sports Medicine
- American Diabetes Association
- American Dietetic Association
- American Heart Association
- American Medical Association
- American Obesity Association
- American Society for Bariatric Surgery
- Built Environments
- Center for Maternal and Child Health
- Center for Nutrition Policy and Promotion
- Center for Science in the Public Interest
- Centers for Disease Control and Prevention
- Child Obesity Programs
- Community Level Initiatives to Prevent Obesity
- Community Programs to Prevent Obesity
- Council on Size and Weight Discrimination
- Economics of Obesity
- Expanded Food and Nutrition Program
- Federal Initiatives to Prevent Obesity
- Food and Drug Administration
- Food Guide Pyramid
- Food Labeling
- Food Marketing to Children
- Food Stamp Nutrition Education Program
- Government Agencies
- Head Start
- Healthy Eating Index
- Healthy People 2010
- National Association to Advance Fat Acceptance
- National Cancer Institute
- National Center for Health Statistics
- National Eating Disorders Association
- National Heart, Lung, and Blood Institute
- National Institutes of Health
- NIDDK
- North American Association for the Study of Obesity
- Obesity in Schools
- Office of Dietary Supplements
- Office of Minority Health
- Policy to Prevent Obesity
- President's Council on Physical Fitness and Sports
- Safety of Urban Environments
- School Initiatives to Prevent Obesity
- Shape-Up America!
- Social Marketing and Obesity
- State and Local Initiatives to Prevent Obesity
- Taxation of Unhealthy Foods
- Toxic Environment
- U.S. Department of Agriculture
- U.S. Department of Health and Social Services
- Weight Control Information Network
- Psychological Influences and Outcomes of Obesity
- Addictive Behaviors
- Anorexia Nervosa
- Anxiety
- Binge Eating
- Bulimia Nervosa
- Cognitive-Behavioral Therapy
- Compulsive Overeating
- Depression
- Disordered Eating
- Eating Disorders in School Children
- External Controls
- Loneliness
- Night Eating Syndrome
- Obsessive Compulsive Disorder
- Psychiatric Medicine and Obesity
- Self-Esteem and Obesity
- Stress
- Suicidality
- Well-Being
- Societal Influences and Outcomes of Obesity
- Alcohol
- Appearance
- Body Image
- Breastfeeding vs. Formula Feeding
- Built Environments
- Calcium Intake and Dairy Products
- Carbohydrate and Protein Intake
- Computers and the Media
- Eating Out in the United States
- Fat Acceptance
- Fat Intake
- Flavor Learning
- Food Advertising and Obesity
- Food Guide Pyramid
- Food Intake Patterns
- Food Labeling
- Food Preferences
- Governmental Policy and Obesity
- Income Level and Obesity
- Nutrition Education
- Obesity and Academic Performance
- Obesity and Drug Use
- Obesity and Sports
- Obesity and the Media
- Obesity in Schools
- Personal Relationships and Obesity
- Physical Activity Patterns in the Obese
- Smoking
- Soda and Soft Drink Intake
- Stereotypes and Obesity
- Supersizing
- Variety of Foods and Obesity
- Virtual Environments
- Weight Discrimination
- Western Diet
- Women and Dieting
- Women and Obesity
- Assessment of Obesity and Health Risks
- Bariatric Surgery in Women
- Body Image
- Breast Cancer
- Breastfeeding
- Colon Cancer
- Coronary Heart Disease in Women
- Early Onset Menarche and Obesity in Women
- Economic Disparities among Obesity in Women
- Endometrial and Uterine Cancers
- Estrogen Levels
- Ethnic Disparities among Obesity in Women
- Exercise and Physical Activity among Obese Women
- Fat Acceptance
- Fertility
- Food Preferences
- Gestational Diabetes
- Implications of Gestational Development
- Maternal Influences on Child Feeding
- Menopause and Obesity
- Morbid Obesity in Women
- Obese Women and Social Stigmatization
- Polycystic Ovary Disease
- Pregnancy Prevalence of Obesity in U.S. Women
- Self-Esteem in Obese Women
- Support Groups for Obese Women
- Waist-to-Hip Ratio
- Women and Diabetes
- Women and Dieting
- Worldwide Prevelance of Obesity
- Loading...
Get a 30 day FREE TRIAL
-
Watch videos from a variety of sources bringing classroom topics to life
-
Read modern, diverse business cases
-
Explore hundreds of books and reference titles

Sage Recommends
We found other relevant content for you on other Sage platforms.

Have you created a personal profile? Login or create a profile so that you can save clips, playlists and searches