Entry
Reader's guide
Entries A-Z
Subject index
Bayesian Networks
A Bayesian network is a graphical representation of a multivariate probability distribution on a set of discrete random variables. Representational efficiency is achieved by explicit separation of information about conditional independence relations between the variables (coded in the network structure) and information about the probabilities involved (coded as a set of numeric parameters or functions). The network structure is expressed as a directed acyclic graph (DAG) that makes the representation amenable to an intuitively appealing, causal interpretation. Algorithms exist for learning both network structure and parameters from data. Furthermore, Bayesian networks allow for computing any marginal or conditional probability regarding the variables involved, thus offering a powerful framework for reasoning with uncertainty. Bayesian networks are also called belief networks and causal probabilistic networks.
Figure 2 A Mixed Treatment Comparison network involving six thrombolytic treatments following acute myocardial infarction and one surgical treatment, percutaneous transluminal angioplasty (PCTA)
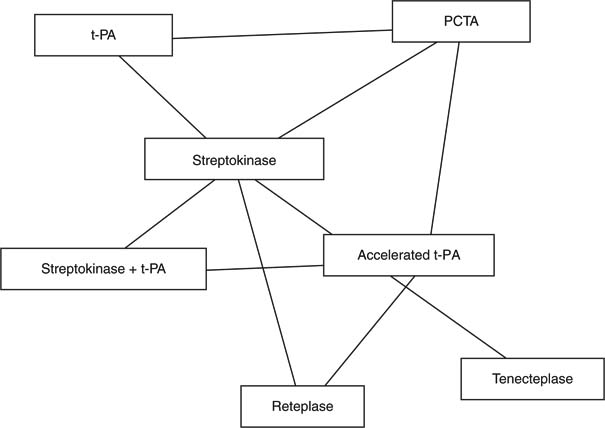
Bayesian networks are suited to model the uncertainty that inheres in many biomedical domains and are, therefore, frequently used in applications of computer-assisted decision making in biomedicine. Furthermore, extensions of Bayesian networks (called influence diagrams) can be used to perform decision analyses.
This entry first sketches the historical background of Bayesian networks. Subsequently, it elaborates on model structure, approaches for network construction, inference methods, medical applications, and software.
Historical Background
Bayesian networks originated in the mid-1980s from the quest for mathematically sound and computationally tractable methods for reasoning with uncertainty in artificial intelligence. In the preceding decade, the first applications of computer-assisted decision making had found their way to the medical field, mostly focusing on the diagnostic process. This had required the development of methods for reasoning with uncertain and incomplete diagnostic information.
One popular method was the naive Bayesian approach that required specification of positive and negative predictive values for each of a set of predefined diagnostic tests and a prior (i.e., marginal) probability distribution over possible diagnostic hypotheses. The approach assumed that all test results were mutually independent markers of disease and used Bayes's theorem to compute posterior (i.e., conditional) probabilities on the hypotheses of interest. The approach is simple and fast and requires a relatively small number of marginal and conditional probabilities to be specified. However, the assumption of independence is mostly wrong and leads to overly extreme posterior probabilities.
Another approach arose in the field of expert systems, where algorithms had been devised to reason with so-called certainty factors, parameters expressing the strength of association in if-then rules. The underlying reasoning principles were mostly ad hoc and not rooted in probability theory, but large sets of if-then rules allowed for a domain representation that was structurally richer and more complex than naive Bayesian models. Bayesian networks bring together the best of both approaches by combining representational expressiveness with mathematical rigor.
Model Structure
Bayesian networks belong to the family of probabilistic graphical models (PGMs), graphs in which nodes represent random variables, and the (lack of) arcs represent conditional independence assumptions. Let G = (V(G), A(G)) be a directed acyclic graph, where the nodes V(G) = {V1, …, Vn} represent discrete random variables with a finite value domain. For each node Vi ∊ V(G), let πi denote the set of parent nodes of Vi in graph G. A Bayesian network now is a pair B = (G, Θ), where Θ={θi|Vi ∊ V(G)} is a set of parametrization functions. The function θi describes a local model for node Vi ∊ V(G) by specifying a conditional probability θi(v|s) for each possible value v of variable Vi and all possible value assignments s to its parents πi. The Bayesian network B defines a unique multivariate probability distribution Pr on V1, …, Vn using the
...
- Basis for Making the Decision
- Acceptability Curves and Confidence Ellipses
- Beneficence
- Bioethics
- Choice Theories
- Construction of Values
- Cost-Benefit Analysis
- Cost-Comparison Analysis
- Cost-Consequence Analysis
- Cost-Effectiveness Analysis
- Cost-Minimization Analysis
- Cost-Utility Analysis
- Decision Quality
- Distributive Justice
- Dominance
- Equity
- Evaluating Consequences
- Expected Utility Theory
- Expected Value of Perfect Information
- Extended Dominance
- Health Production Function
- League Tables for Incremental Cost-Effectivenes: Ratios
- Marginal or Incremental Analysis, Cost-Effectiveness Ratio
- Monetary Value
- Moral Choice and Public Policy
- Net Benefit Regression
- Net Monetary Benefit
- Nonexpected Utility Theories
- Pharmacoeconomics
- Protected Values
- Rank-Dependent Utility Theory
- Return on Investment
- Risk-Benefit Trade-Off
- Subjective Expected Utility Theory
- Toss-Ups and Close Calls
- Value-Based Insurance Design
- Welfare, Welfarism, and Extrawelfarism
- Biostatistics and Clinical Epidemiology
- Analysis of Covariance (ANCOVA)
- Analysis of Variance (ANOVA)
- Attributable Risk
- Basic Common Statistical Tests: Chi-Square Test, t Test, Nonparametric Test
- Bayes's Theorem
- Bayesian Analysis
- Bayesian Evidence Synthesis
- Bayesian Networks
- Bias
- Bias in Scientific Studies
- Brier Scores
- Calibration
- Case Control
- Causal Inference and Diagrams
- Causal Inference in Medical Decision Making
- Conditional Independence
- Conditional Probability
- Confidence Intervals
- Confounding and Effect Modulation
- Cox Proportional Hazards Regression
- Decision Rules
- Diagnostic Tests
- Discrimination
- Distributions: Overview
- Dynamic Treatment Regimens
- Effect Size
- Equivalence Testing
- Experimental Designs
- Factor Analysis and Principal Components Analysis
- Fixed Versus Random Effects
- Frequentist Approach
- Hazard Ratio
- Hypothesis Testing
- Index Test
- Intraclass Correlation Coefficient
- Likelihood Ratio
- Log-Rank Test
- Logic Regression
- Logistic Regression
- Maximum Likelihood Estimation Methods
- Measures of Central Tendency
- Measures of Frequency and Summary
- Measures of Variability
- Meta-Analysis and Literature Review
- Mixed and Indirect Comparisons
- Multivariate Analysis of Variance (MANOVA)
- Nomograms
- Number Needed to Treat
- Odds and Odds Ratio, Risk Ratio
- Ordinary Least Squares Regression
- Parametric Survival Analysis
- Poisson and Negative Binomial Regression
- Positivity Criterion and Cutoff Values
- Prediction Rules and Modeling
- Probability
- Propensity Scores
- Randomized Clinical Trials
- Receiver Operating Characteristic (ROC) Curve
- Recurrent Events
- Recursive Partitioning
- Regression to the Mean
- Sample Size and Power
- Screening Programs
- Statistical Notations
- Statistical Testing: Overview
- Subjective Probability
- Subset Analysis: Insights and Pitfalls
- Survival Analysis
- Tables, Two-by-Two and Contingency
- Variance and Covariance
- Violations of Probability Theory
- Weighted Least Squares
- Decision Analysis and Related Mathematical Models
- Applied Decision Analysis
- Boolean Algebra and Nodes
- Decision Analyses, Common Errors Made in Conducting
- Decision Curve Analysis
- Decision Tree: Introduction
- Decision Trees, Advanced Techniques in Constructing
- Decision Trees, Construction
- Decision Trees, Evaluation
- Decision Trees, Evaluation With Monte Carlo
- Decision Trees: Sensitivity Analysis, Basic and Probabilistic
- Decision Trees: Sensitivity Analysis, Deterministic
- Declining Exponential Approximation of Life Expectancy
- Deterministic Analysis
- Discrete-Event Simulation
- Disease Management Simulation Modeling
- Expected Value of Sample Information, Net Benefit of Sampling
- Influence Diagrams
- Markov Models
- Markov Models, Applications to Medical Decision Making
- Markov Models, Cycles
- Markov Processes
- Reference Case
- Steady-State Models
- Stochastic Medical Informatics
- Subtrees, Use in Constructing Decision Trees
- Test-Treatment Threshold
- Time Horizon
- Tornado Diagram
- Tree Structure, Advanced Techniques
- Health Outcomes and Measurement
- Complications or Adverse Effects of Treatment
- Cost-Identification Analysis
- Costs, Direct Versus Indirect
- Costs, Fixed Versus Variable
- Costs, Opportunity
- Costs, Out-of-Pocket
- Costs, Semifixed Versus Semivariable
- Costs, Spillover
- Economics, Health Economics
- Efficacy Versus Effectiveness
- Efficient Frontier
- Health Outcomes Assessment
- Health Status Measurement Standards
- Health Status Measurement, Assessing Meaningful Change
- Health Status Measurement, Construct Validity
- Health Status Measurement, Face and Content Validity
- Health Status Measurement, Floor and Ceiling Effects
- Health Status Measurement, Generic Versus Condition-Specific Measures
- Health Status Measurement, Minimal Clinically Significant Differences, and Anchor Versus Distribution Methods
- Health Status Measurement, Reliability and Internal Consistency
- Health Status Measurement, Responsiveness and Sensitivity to Change
- Human Capital Approach
- Life Expectancy
- Morbidity
- Mortality
- Oncology Health-Related Quality of Life Assessment
- Outcomes Research
- Patient Satisfaction
- Regret
- Report Cards, Hospitals and Physicians
- Risk Adjustment of Outcomes
- SF-36 and SF-12 Health Surveys
- SF-6D
- Sickness Impact Profile
- Sunk Costs
- Impact or Weight or Utility of the Possible Outcomes
- Certainty Equivalent
- Chained Gamble
- Conjoint Analysis
- Contingent Valuation
- Cost Measurement Methods
- Decomposed Measurement
- Disability-Adjusted Life Years (DALYs)
- Discounting
- Discrete Choice
- Disutility
- EuroQol (EQ-5D)
- Health Utilities Index Mark 2 and 3 (HUI2, HUI3)
- Healthy Years Equivalents
- Holistic Measurement
- Multi-Attribute Utility Theory
- Person Trade-Off
- Quality of Well-Being Scale
- Quality-Adjusted Life Years (QALYs)
- Quality-Adjusted Time Without Symptoms or Toxicity (Q-TWiST)
- SMARTS and SMARTER
- Split Choice
- Utilities for Joint Health States
- Utility Assessment Techniques
- Willingness to Pay
- Other Techniques, Theories, and Tools
- Artificial Neural Networks
- Bayesian Networks
- Bioinformatics
- Chaos Theory
- Clinical Algorithms and Practice Guidelines
- Complexity
- Computer-Assisted Decision Making
- Constraint Theory
- Decision Board
- Decisional Conflict
- Error and Human Factors Analyses
- Ethnographic Methods
- Expert Systems
- Patient Decision Aids
- Qualitative Methods
- Story-Based Decision Making
- Support Vector Machines
- Team Dynamics and Group Decision Making
- Threshold Technique
- Perspective of the Decision Maker
- Advance Directives and End-of-Life Decision Making
- Consumer-Directed Health Plans
- Cultural Issues
- Data Quality
- Decision Making in Advanced Disease
- Decisions Faced by Hospital Ethics Committees
- Decisions Faced by Institutional Review Boards
- Decisions Faced by Nongovernment Payers of Healthcare: Managed Care
- Decisions Faced by Patients: Primary Care
- Decisions Faced by Surrogates or Proxies for the Patient, Durable Power of Attorney
- Diagnostic Process, Making a Diagnosis
- Differential Diagnosis
- Evaluating and Integrating Research Into Clinical Practice
- Evidence Synthesis
- Evidence-Based Medicine
- Expert Opinion
- Genetic Testing
- Government Perspective, General Healthcare
- Government Perspective, Informed Policy Choice
- Government Perspective, Public Health Issues
- Health Insurance Portability and Accountability Act Privacy Rule
- Health Risk Management
- Informed Consent
- Informed Decision Making
- International Differences in Healthcare Systems
- Law and Court Decision Making
- Medicaid
- Medical Decisions and Ethics in the Military Context
- Medical Errors and Errors in Healthcare Delivery
- Medicare
- Models of Physician–Patient Relationship
- Patient Rights
- Physician Estimates of Prognosis
- Rationing
- Religious Factors
- Shared Decision Making
- Surrogate Decision Making
- Teaching Diagnostic Clinical Reasoning
- Technology Assessments
- Terminating Treatment, Physician Perspective
- Treatment Choices
- Trust in Healthcare
- The Psychology Underlying Decision Making
- Accountability
- Allais Paradox
- Associative Thinking
- Attention Limits
- Attraction Effect
- Automatic Thinking
- Axioms
- Biases in Human Prediction
- Bounded Rationality and Emotions
- Certainty Effect
- Cognitive Psychology and Processes
- Coincidence
- Computational Limitations
- Confirmation Bias
- Conflicts of Interest and Evidence-Based Clinical Medicine
- Conjunction Probability Error
- Context Effects
- Contextual Error
- Counterfactual Thinking
- Cues
- Decision Making and Affect
- Decision Modes
- Decision Psychology
- Decision Weights
- Decision-Making Competence, Aging and Mental Status
- Deliberation and Choice Processes
- Developmental Theories
- Dual-Process Theory
- Dynamic Decision Making
- Editing, Segregation of Prospects
- Emotion and Choice
- Errors in Clinical Reasoning
- Experience and Evaluations
- Fear
- Frequency Estimation
- Fuzzy-Trace Theory
- Gain/Loss Framing Effects
- Gambles
- Hedonic Prediction and Relativism
- Heuristics
- Human Cognitive Systems
- Information Integration Theory
- Intuition Versus Analysis
- Irrational Persistence in Belief
- Judgment
- Judgment Modes
- Learning and Memory in Medical Training
- Lens Model
- Lottery
- Managing Variability and Uncertainty
- Memory Reconstruction
- Mental Accounting
- Minerva-DM
- Mood Effects
- Moral Factors
- Motivation
- Numeracy
- Overinclusive Thinking
- Pain
- Pattern Recognition
- Personality, Choices
- Preference Reversals
- Probability Errors
- Probability, Verbal Expressions of
- Problem Solving
- Procedural Invariance and Its Violations
- Prospect Theory
- Range-Frequency Theory
- Risk Attitude
- Risk Aversion
- Risk Communication
- Risk Perception
- Scaling
- Social Factors
- Social Judgment Theory
- Stigma Susceptibility
- Support Theory
- Uncertainty in Medical Decisions
- Unreliability of Memory
- Value Functions in Domains of Gains and Losses
- Worldviews
- Loading...
Get a 30 day FREE TRIAL
-
Watch videos from a variety of sources bringing classroom topics to life
-
Read modern, diverse business cases
-
Explore hundreds of books and reference titles

Sage Recommends
We found other relevant content for you on other Sage platforms.

Have you created a personal profile? Login or create a profile so that you can save clips, playlists and searches